Proteomics applications in medicine, basic biology, and beyond

Tyler Ford
September 15, 2022
The proteins that make up the proteome are fundamental to life. So, it’s no surprise applications of proteomics extend to nearly every domain of biology. With new proteomic insights from next-generation proteomic analysis tools, we hope to gain mechanistic insights into how the body works in health and disease, discover new biomarkers, develop new drugs, and far more.
For example, researchers are applying proteomics to better understand the tau and amyloid-beta proteins implicated in the progression of Alzheimer’s disease, while other scientists are using proteomics to learn new things about the plant-microbe interactions that keep crops around the world healthy. These proteomics applications and more are enabled by a growing set of tools and techniques for studying the proteome that are constantly expanding the boundaries of our proteomic knowledge.
Nautilus is excited to be a leader in the proteomics revolution with the potential to enable research breakthroughs through the NautilusTMTM Proteome Analysis Platform. In this blog post, we cover some of the many ways proteomics can drive new discoveries through applications across a wide variety of fields. We cover many of these applications in more depth in other applications of proteomics blog posts and encourage you to subscribe to learn more about the potential of proteomics.
What is proteomics
Proteomics is the study of all the proteins in a biological sample, for example a cell, tissue or organism. These proteins are known collectively as the proteome, carry out the basic functions of cells, and are foundational to most all of biology.
A single cell contains billions of proteins, and it is their dynamic make-up and interactions that determine cellular behavior. Instead of studying individual proteins in isolation, proteomics research aims to study all proteins and their interactions, at the same time.
Proteomics techniques
Recognizing the potential insights that can come from studying the proteome, many researchers have made forays into studying it at scale. They’ve developed proteomics techniques and technologies aiming to provide a comprehensive view of the proteome in any sample of interest. These techniques include antibodies and other affinity reagents, mass spectrometry, and protein sequencing. Unfortunately, traditional proteomics technologies have only been able to scratch the surface of the full proteome and have not delivered on the promise of this exciting field.
Next-generation proteomics platforms are finally poised to deliver on the promise of the proteome thanks to advances in a variety of enabling technologies. We’re moving beyond the scale of traditional technologies and to proteomic analysis platforms that can capture billions of proteins, identify them, and measure their abundance across a wide dynamic range. With advances in nanofabrication, machine learning, data storage, and more, we’re designing technologies with the potential to analyze nearly all the proteins in the human proteome at once. These proteomics technologies will drive great advances in healthcare, basic research, and much more.
Applications of proteomics
The Nautilus Proteome Analysis Platform is designed to measure the majority of the proteome and provide information-rich but easy-to-understand protein analysis. We aim for our platform to provide researchers with information on >95% of the proteome of any sample. With this depth of coverage, we hope researchers can use our platform to expand upon existing proteomic applications and even create entirely new uses for proteomics. We discuss some of the many exciting applications of proteomic data below.
Learn how Kathryn Lilley and others are using proteomics to learn more about protein function.
Proteomics applications in medicine
Prior to observing signs of disease at the whole-organism level, it is likely that the proteome of affected tissues will begin to change. Fluctuations in the proteome can act as biomarkers that predict how patients’ health will change or how they will respond to a treatment. Thus, one application of proteomics is to forecast changes in health by enabling physicians to analyze the proteomes of patient samples. With advanced knowledge of biological changes at the molecular level, physicians may be able to prevent health issues or lower their severity by prescribing preventative measures.
For instance, a physician might begin to see proteins associated with heart disease increase in a patient during a routine check-up. To prevent heart issues, the physician might put this patient on drugs that lower cholesterol.
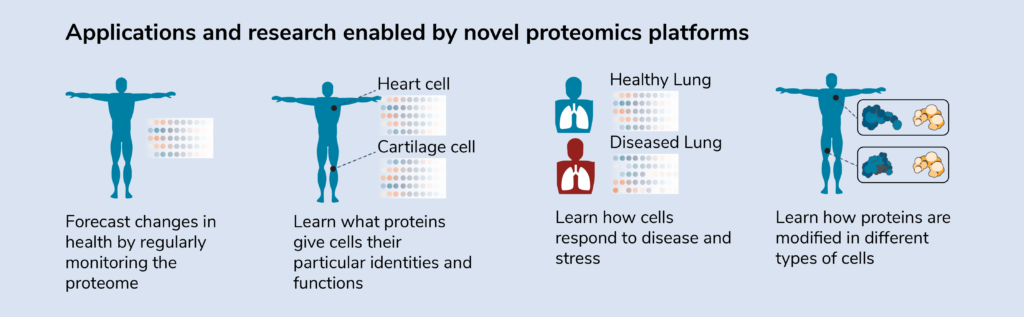
Proteomics applications in disease research
Although researchers know the symptoms of many diseases and may know what genes or pathogens drive them, they do not necessarily have a good understanding of the underlying molecular mechanisms. That is, they don’t always know how diseases alter proteins to change cellular function. Proteomics can reveal how the composition and abundance of proteins change when cells are diseased or stressed, enabling researchers to create treatments that counter such changes.
For example, in a disease like COVID-19, researchers might see that patient samples have an overabundance of immune system proteins. They could then prescribe these patients drugs that tamp down the effects of the immune-activating proteins to protect patients from autoimmune damage.
Researchers have also applied proteomics in numerous scientific studies examining the roots of Alzheimer’s disease. For example, researchers used a targeted proteomics approach to examine the ways the tau protein is modified in Alzheimer’s disease, finding a number of tau proteoforms linked to the progression of Alzheimer’s. In another proteomics application, scientists combined proteomics and genomics in a multiomic approach that uncovered hundreds of regions of DNA correlated with proteins that affected Alzheimer’s risk.
Proteomics applications in cancer research have proven extraordinarily useful as well. One recent paper provided new and potentially more accurate subtypes for breast cancer based on a discovery proteomic analysis of samples from patients. Those subtypes could in turn allow doctors to design more precise treatments for patients. Another study including a proteomic analysis identified both potential biomarkers and therapeutic targets in pancreatic cancer, such as specific glycoproteins.
- Learn more about applications of proteomics in cancer research
- Learn more about applications of proteomics in neuroscience
Listen to the Translating Proteomics podcast to learn how proteomics can help expand the drug development toolkit.
Proteomics applications in basic research – Understanding cellular identity and function
The genome of an organism is – with some exceptions – the same in every cell, but the proteome of different cells within an individual varies widely. By profiling the protein composition of different cell types, proteomics researchers can learn what proteins give cells their functions.
Eventually, they may even be able to manipulate the proteome of one cell to turn it into another kind of cell. For instance, using insights from proteomics physicians may one day be able to generate replacement cells to heal damaged tissues or organs.
Check out this episode of the “Translating Proteomics” podcast to learn how we can leverage proteomics to study “Biology in Space and Time”
Applications of proteomics in agriculture
Proteomics has a wide range of potential uses in agriculture. For example, farmers could use proteomics to forecast changes in crop health. Before entire crops begin to die off, farmers might be able to check them for changes in protein biomarkers associated with viral or bacterial infections. Then they could apply protective chemicals throughout the farm and prevent the spread of the disease, improving crop yields.
Proteomics could also find applications when it comes to fertilizing crops. Humanity currently spends a vast amount of energy and money creating nitrogen-based fertilizers. Interestingly, there are plants that naturally cooperate with bacteria in the soil and create nitrogenous nutrients from the abundant nitrogen in the air, converting it into a form plants can use. Researchers could measure the proteomes of these nitrogen-fixing bacteria and the plant cells that interact with them to better understand how they cooperate to facilitate this transfer of nutrients.
With a better understanding of how the proteome enables this process, researchers may be able to give other plants the ability to create their own nitrogenous nutrients and thereby make it possible to grow more food for a growing global population using less fertilizer.
Proteomic applications in agriculture could also extend to safeguarding environmental health as the planet faces threats from climate change. For instance, scientists will be able to use proteomics to study how various plants’ proteomes respond to climatic stresses like drought. If they find plants that fare well in droughts, they can compare their proteomes to those of plants that die off during drought. They may find proteins or sets of proteins that make plants particularly drought resistant. Researchers might then increase the levels of these resistance proteins in drought-susceptible plants, helping them to survive. This could help preserve ecosystem health as the climate changes.
Applications of proteomics – paleoproteomics
Proteomics is finding applications in the world of archaeology and anthropology as well. Proteins can survive for far longer than DNA can in many circumstances, allowing scientists to glean important information about ancient people and animals that would otherwise be lost.
Examples of paleoproteomic applications include using proteins from teeth to show what people ate thousands of years ago, reconstructing ancient trade routes using proteins from different plants, and even revealing the proteomes of extinct creatures.
New horizons in proteomics research
We are only barely skimming the surface of what proteomics can do for applied and basic research across many different fields. Nonetheless, we hope you are convinced that the potential of novel proteomics technologies is vast and largely untapped. In other blog posts, we dive into more detail on how scientists study the proteome and take a look at the types of biological information they can uncover using various proteomics methods.
Find more applications of proteomics.
Check out the Translating Proteomics podcast for an exciting discussion of applications of proteomics
MORE ARTICLES

