Targeting healthcare disparities through the proteomics revolution (Part 1)

Tyler Ford
January 17, 2023
Proteins generally determine whether people are healthy or sick regardless of their genomic background. Thus, comprehensively mapping the precise makeup and distribution of proteins in the body, the “proteome,” will greatly advance researchers’ and physicians’ knowledge of health and disease. With an improved understanding of the proteome, researchers will be better able to design novel treatments that target diseases at their mechanistic roots in proteins and monitor treatment effectiveness through protein fluctuations.
Next generation proteomics technologies, like the Nautilus Proteome Analysis Platform, aim to give researchers the ability to quickly and comprehensively analyze the proteome. These technologies will hopefully help us achieve a healthier future. But a critical question is, for whom? To ensure the benefits of proteomics reach all people, we need to make the data and applications derived from these technologies available to everyone regardless of background, ethnicity, geographic location, or any other factors.
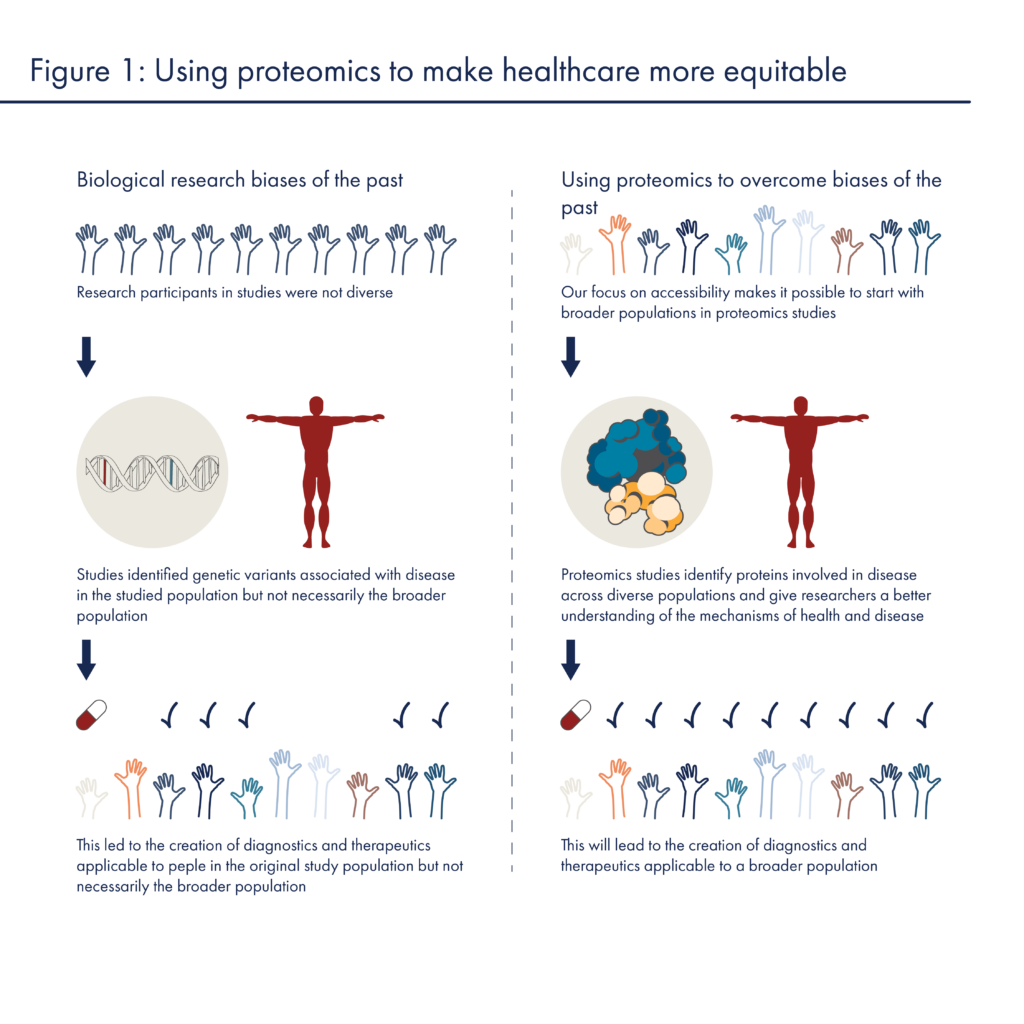
In the past, biological research efforts have been marred by fundamental biases that limit the scope of their findings as well as their utility. To combat these biases, many, including the American Association for Cancer Research and the National Academies of Sciences Engineering and Medicine, have called for concerted efforts to improve representation in biological and clinical research. As proteomics begins to mature, we and others have the opportunity to avoid the biases of the past and create a more equitable healthcare future for all.
This post describes the ways biases have historically led to inequities in genomics research and highlights ways researchers can rectify those inequalities with more and better data. In part 2 of this series, we will discuss how novel proteomics technologies may enable a more equitable future.

Equity downfalls in the genomics era
To understand how research biases have undermined health equity in the past, it is informative to look to genomics. It is an understatement to say that much good has come from sequencing the human genome. With a first (and quite rough) draft of the human genome completed in the early 2000s, researchers gained access to information and technologies that created a scaffold upon which they could build hypotheses about the causes of a wide variety of diseases. They began to quickly associate many specific genetic variants with diseases. Unfortunately, this scaffolding was built off of genome sequences that came almost exclusively from people of European descent. There was very limited representation of people from other ancestries and, even today 87% of people in large scale genomic studies are of European descent. This limits the benefits to people from other ancestries in a number of ways:
- Genetic variants associated with diseases in people from one racial, ethnic, or geographic background may not be associated with those diseases in other backgrounds. Often, when researchers find genetic variants associated with diseases, the mechanisms through which they cause disease are unknown. In many cases, the associated variants don’t cause the disease at all, but may just be very close to other regions of the genome that encode broken proteins or cause disease in some other way. The identified variants are associated with disease but it cannot be assumed that they cause disease or that they will be associated with the disease across all populations. By including more diverse populations in genomics studies, it is both possible to identify associations that are specific to people from different backgrounds and to increase the potential to find truly causal variants.
- Interactions between genetic variants in different populations may mitigate or exacerbate disease. Even if a genetic variant encodes a protein that causes a disease in one population, that does not mean that it will cause that disease in another population. For instance, one group of people may harbor compensatory variants elsewhere in the genome that prevent the original “disease” variant from being a problem. If genomics studies include people from many different backgrounds, they may show that a disease fails to manifest even in the face of a supposedly causal variant. This provides opportunities for future studies to dive into the genetics of the unaffected population and learn more about the biology of the disease.
- Environmental factors specific to certain subpopulations may interact with DNA variants to cause disease. For instance, if one subpopulation eats a plant that is turned into a toxin by a protein encoded in a genetic variant, diseases caused by that toxin will only appear in the subpopulation that eats the plant. Similarly, some drugs may be toxic to people with specific genetic variants but perfectly fine in people without those variants. Again, unless people from diverse groups are included in the studies relating genetics to such diseases and drugs, it will be difficult to understand the mechanisms behind their harmful effects.
(For a more thorough look at these and other issues of equity in genomics see Sirugo et al 2019.)
These issues lead to cascading inequities in healthcare. With biased genomics data, researchers create tests that only diagnose patients from particular subpopulations and treatments that are only effective in these subpopulations. For example, most screens for cystic fibrosis only identify genetic variants commonly found in people of European descent (Lim et al 2016), and many people with uncommon variants are ineligible for recently approved treatments (McGarry and McColley 2021). These inequities lead to mistrust in the biomedical system and deter minority participation in health research (George et al 2014).
With the proteomics revolution, we have the chance to avoid the biases of the past and engender trust in an improved healthcare system. We’ll discuss how in the next post in this series.
MORE ARTICLES
