3 ways next-generation proteomics can impact neuroscience – Applications of proteomics in neuroscience

Tyler Ford
July 13, 2023
Neuroscience research has advanced dramatically over the last few decades. Researchers now have the ability to trace neuronal connections, activate brain pathways using optogenetics, and even grow mini brains in the lab. Nonetheless, we still have limited understanding of how the brain and broader nervous system work in healthy individuals and malfunction in disease.
Proteomics offers the ability to measure the protein composition of tissues, cells, and specialized cellular structures in the nervous system. This could point researchers to proteins essential for the neurological processes involved in learning, memory, and a wide variety of behaviors. It could also help them identify biomarkers and drug targets for neurological disorders.
While traditional proteomics technologies often fail to measure the full proteome and can be blind to the precise patterns of modification found on proteoforms that drive changes in cellular activities, next-generation proteomics platforms like the Nautilus Proteome Analysis Platform are designed to measure substantively all of the proteome in any cell type.
What is next-generation proteomics?
Next-generation proteomics refers to up-and-coming proteomics technologies designed to enable researchers to fully analyze and understand the proteome. Current proteomics technologies like mass spectrometry cannot routinely provide comprehensive proteome coverage. Next-generation proteomics tools like the Nautilus Platform are designed to be accessible to more researchers and have the sensitivity and dynamic range to identify every protein in a sample, making true exploration of the proteome possible.
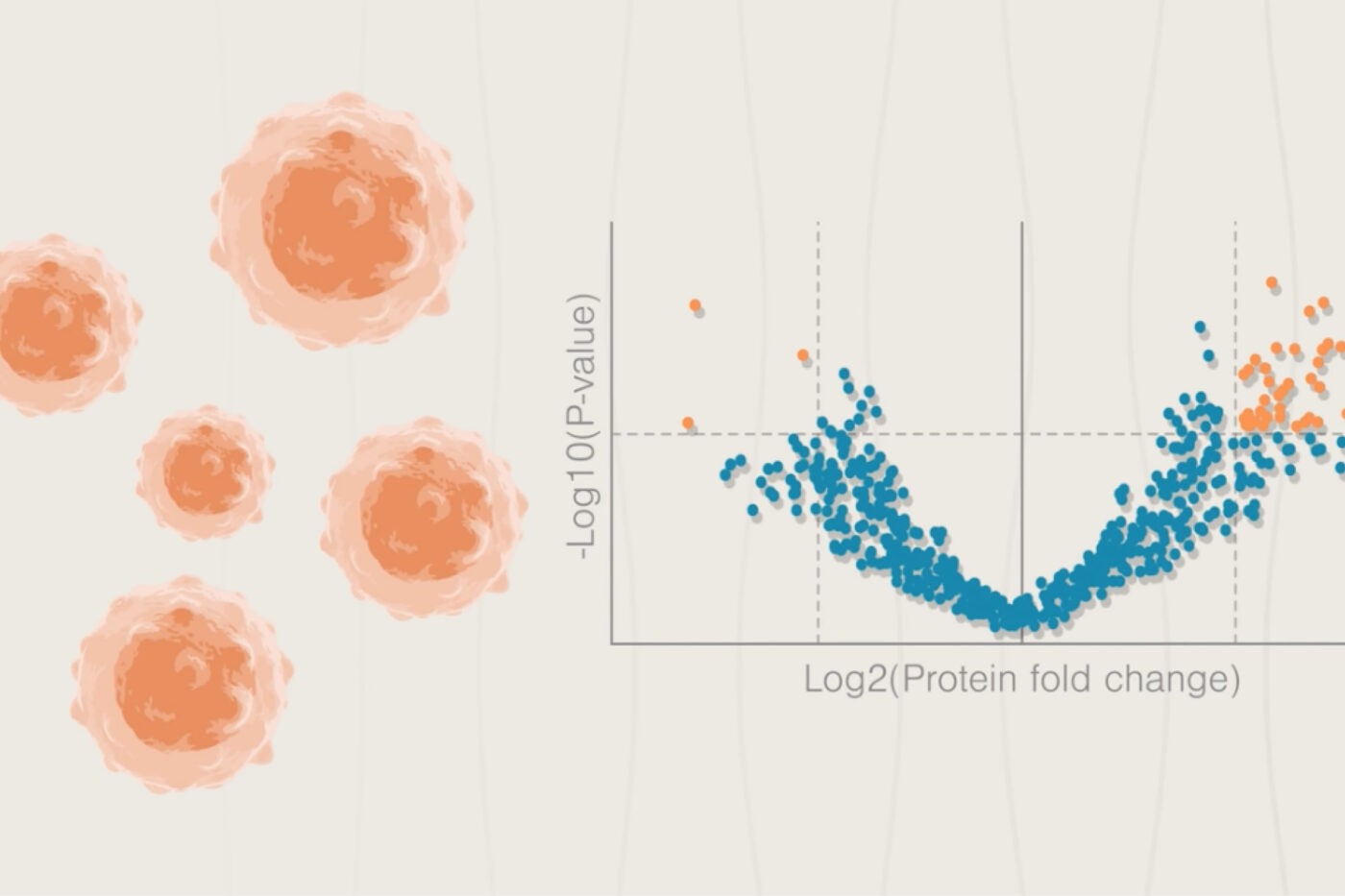
Here, we discuss three ways next-generation proteomics platforms can enable neuroscience research. This is far from an exhaustive list, but demonstrates how next-generation proteomics can transform neuroscience.
Watch our animation to see how next-generation proteomics can fuel neuroscience research
Advancing our understanding of neuronal development with proteomics
The brain is an incredibly complex and heterogeneous organ and understanding how the architecture of the brain and broader nervous system comes to be is a key goal of neuroscience. While scientists have learned a great deal about the cues that establish and maintain neuronal connections, there is much to learn about the molecular mechanisms that guide these connections.
Scientists are using proteomics to learn what proteins are altered as a result of signaling in neuronal development. Techniques such as interaction proteomics can show where these proteins localize and clue researchers into their functions. Techniques like phosphoproteomics, on the other hand, can reveal signaling pathways active during various developmental stages and help reveal the mechanisms behind neural plasticity.
Proteoform profiling is particularly important for studying neuronal development. Things like cytoskeletal proteins are dynamically regulated in neuronal development and regeneration to create and maintain the extensive cellular contacts in the nervous system. The stability of the cytoskeleton is partially determined by posttranslational modifications to cytoskeletal proteins and understanding the composition, distribution, and abundance of these cytoskeletal and other proteoforms in various neuronal contexts will be essential for a thorough understanding of the nervous system.
Next-generation proteomics platforms are expected to make it much easier to identify all the players involved in neuronal development including their various proteoforms. This may enable scientists to gain a more thorough understanding of things like cytoskeletal modifications that are essential to the dynamics of the nervous system.
Advancing our understanding of neuronal cell signaling with next-generation proteomics
Neurons transmit electrochemical signals to store, process, and relay information as well as to coordinate activities with other cells and tissues. Neuronal networks involved in these functions can be extremely complicated. For example, there are billions of interconnected neurons in the mammalian brain. Researchers have begun mapping these networks and have even mapped full neuronal networks in animals like C. elegans, but one key to understanding their connections and functions is to learn more about the types of signals sent between neurons and how different kinds of neurons receive and respond to those signals.
Synapses are specialized structures at neuronal interfaces where interneuronal communication is largely defined. Here, neurotransmitters are released by pre-synaptic neurons and received by post-synaptic neurons. These neurotransmitters bind to receptors in the post-synaptic neurons and act through various signaling cascades and protein channels to either activate or inhibit the post-synaptic neuron. Understanding what machinery is involved in both the release and receipt of neurotransmitters (of which there are many) is essential to understanding nervous system function.
Researchers have developed a variety of techniques to measure the proteomes of neurons in discrete areas of the brain and more specifically in synapses. In one such technique, researchers genetically encode labeling enzymes such that they are produced in particular regions of the brain, particular types of neurons, and eventrafficked to the synapses. These enzymes then add labels to proteins found in these specific regions and the labels can be used to isolate the proteins. Currently, researchers use mass spectrometry to identify such labeled proteins. While useful, mass spectrometry has sensitivity issues and usually cannot identify specific proteoforms. This can make it difficult to get a full picture of the proteomes of specific neurons or synapses. For a review on these techniques, see Wang and Savas 2018.
Next-generation proteomics platforms like the Nautilus Proteome Analysis Platform are designed to have single-molecule sensitivity. The Nautilus Platform also analyzes intact proteins and can therefore identify different proteoforms with relative ease. These technologies should make it easier to understand the precise proteomic make-up of particular neurons and synapses and thereby provide researchers with a mechanistic understanding of the neuronal signaling pathways involved in various behaviors and nervous system activities.
Advancing our understanding of neurological disease with next-generation proteomics
Perhaps the best-known neurological disorder is Alzheimer’s disease. More than 6 million people in the US alone have this progressive, debilitating, and ultimately lethal disease. It’s expected that this number will grow dramatically over the next few decades as the American population ages.
While the precise molecular events causing this disease are not completely understood, Alzheimer’s pathology has been associated with the aggregation:
- Amyloid beta outside brain cells
- Tau inside brain cells
Scientists have already begun developing drugs designed to remove these aggregates and hopefully slow or stop progression of the disease. While there has been moderate success, there is still much to learn about what causes these proteins to aggregate and how they’re involved in neurological dysfunction. In fact, some drugs have effectively removed aggregates without abrogating disease while others slow but do not stop disease progression. Clearly more research is needed.
Recent work with traditional proteomics platforms has begun to reveal the role post translational modifications play in the development of aggregates and Alzheimer’s disease. For example, Wesseling, et al showed that various phosphorylated, ubiquitinated, and acetylated forms of the tau protein correlate with clinical stages of Alzheimer’s disease. This work points to the need to target different forms of tau at different stages of the disease.
Such studies begin to elucidate tau’s role in disease biology, but are difficult to perform on standard proteomics platforms. These technologies are not routinely used to identify proteoforms in many labs. Next-generation proteomics platforms aim to routinely measure the collection of proteoforms in cells. Such studies should provide researchers with an in-depth view of protein modification in Alzheimer’s and many other diseases involving protein modification such as Parkinson’s and prion diseases. These studies may lead to the development of novel treatments targeting pathogenic proteoforms and their effects.
In addition to elucidating the mechanisms underlying diseases like Alzheimer’s, next-generation proteomics can also help researchers develop novel biomarkers for these diseases. After cognitive defects are identified, Alzheimer’s is currently diagnosed through brain imaging and the measurement of Alzheimer’s proteins in cerebral spinal fluid. Blood tests are also in development, but novel tests that are less invasive and help doctors better identify the stage of the disease as well as potential treatments are needed. Efforts are underway in labs like that of Randall J. Bateman to identify new Alzheimer’s biomarkers using mass spectrometry. Their efforts may lead to the development of more effective diagnostics and treatments.
Nonetheless, most traditional proteomics platforms lack the sensitivity and dynamic range to assess such biomarkers routinely. Next-generation platforms are designed to measure the majority of the proteome even in a sample as complicated as blood plasma. This may make it possible to treat these diseases early, before neurological impacts become pronounced.
We want to work with you to enable neuroscience research with the power of next-generation proteomics
We’re just scratching the surface in terms of the many ways next-generation proteomics platforms can enhance our understanding of neuroscience. Although this field is complex, the single-molecule analysis capabilities and high dynamic range of technologies like the Nautilus Proteome Analysis Platform are designed to be up to the challenge. The Nautilus Platform in particular is designed to interrogate intact, full-length proteins at the single molecule level and is expected to be able to resolve the patterns of modification on individual proteins in a sample of interest through proteoform analyses. If you’re working in this field and would like to collaborate to transform neuroscience, please reach out!
Check out our round up of proteomics and neuroscience posts.
MORE ARTICLES

