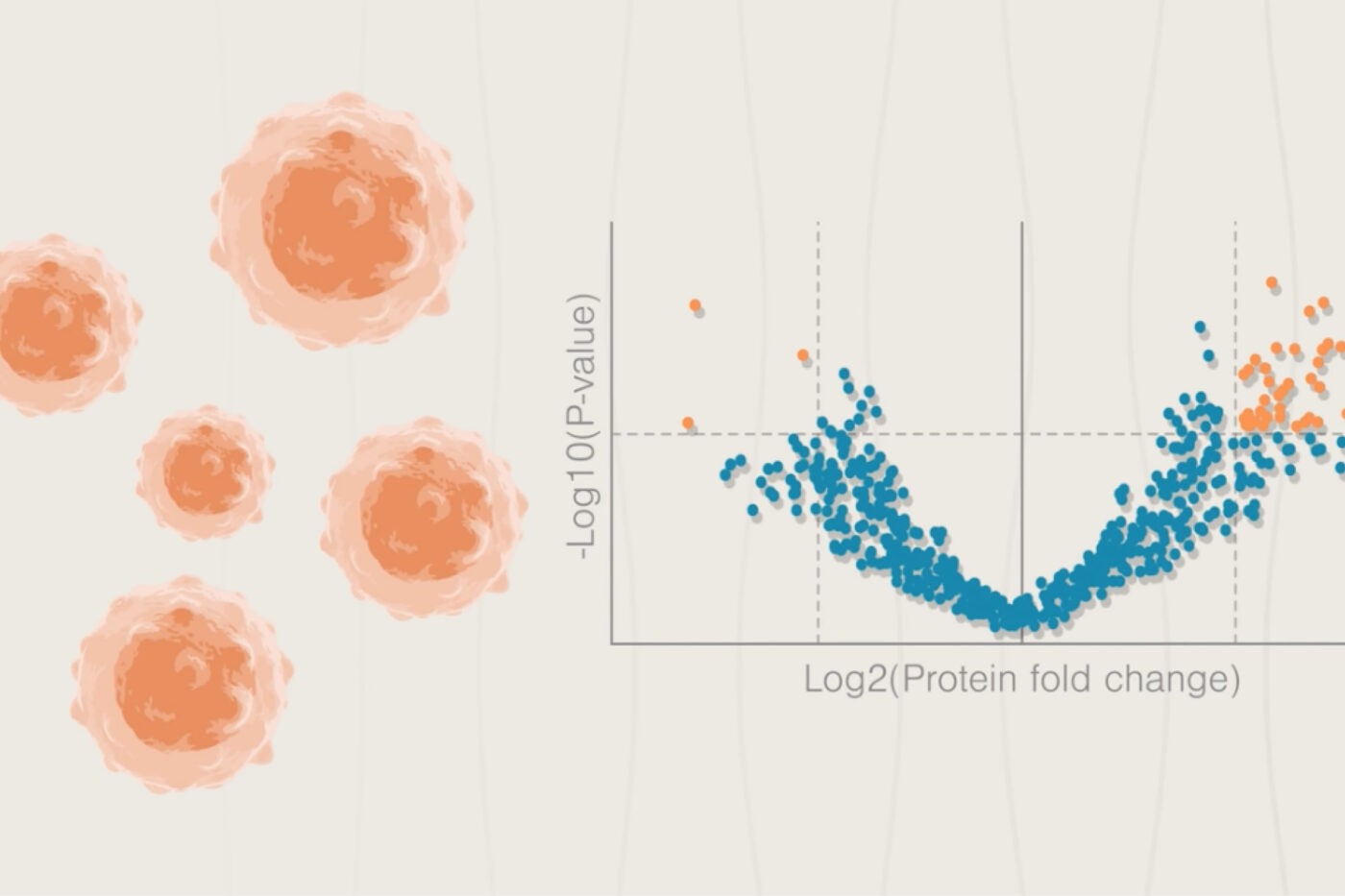
Proteomics is playing a pivotal role in the field of cancer research, enabling scientists to gain new insights into fundamental cancer biology, identify new biomarkers, and much more. Recent research makes it plain that genomics is powerful, but insufficient for understanding how cancers survive and grow. Adding proteomics and other elements of multiomics to the cancer research toolkit yields rich insights by capturing details at different levels of biology that genomic analyses miss. For example, multiomics studies can follow the effects of a cancer driver mutation through the levels of the central dogma and point to numerous potential drug targets.
With next-generation proteomics tools, efforts to understand and ultimately defeat many different kinds of cancer will hopefully accelerate. Using proteomics tools that are more comprehensive , easier to use, and higher throughput, scientists will be able to link more proteins to cancer outcomes, tie them to underlying genetic mutations, and discover how they disrupt normal biological processes. With easier workflows, more labs will be able to conduct this kind of research, further broadening the reach of the field.
Here, we compile content that highlights how cancer researchers can leverage proteomic analyses to make new discoveries and push the field forward. Click through the links below to find forward looking content focused on the potential of proteomics and in-depth summaries of recent work at the intersection of proteomics and cancer.
Find our full proteomics and cancer ebook here.
Proteomics applications in the development of precision medicines against cancer
Developing cancer drugs that target specific genetic mutations or aspects of a patient’s particular cancer are primary goals of precision medicine. This is harder than it sounds, however. Finding the right genetic mutations is difficult, and identifying the proteins impacted by those mutations can also pose a challenge. The field of proteomics, including up-and-coming next-generation proteomics tools, is poised to accelerate this effort by enabling researchers to find proteins associated with cancer more quickly and comprehensively. Our series on precision medicine and proteomics covers this topic in detail.
Watch our animation to see how next-generation proteomics can fuel cancer research:
Multiomics analysis uncovers biological pathways at work in lung cancer
Gillette et al. used a multiomic analysis to investigate more than 100 samples from lung adenocarcinoma patients. They gleaned insights into new lung cancer subtypes, the effects of kinase fusions, and mutations leading to immune evasion. Their work also identified protein biomarkers associated with druggable mutations and highlighted disparities between mRNA and protein abundance, underscoring the need for proteomic analyses of complex diseases like lung adenocarcinoma.
Proteomic analysis aids cancer metastasis investigation
The epithelial to mesenchymal transition (EMT) is an important process in cancer metastasis, allowing tumor cells to move throughout the body and grow elsewhere. Paul et al. provide important new insights into this process using a proteomic analysis along with other techniques to watch this process play out in vitro.
One key finding from their work was that data across different omes was poorly correlated, meaning multiomic analyses are necessary to fully understand the epithelial to mesenchymal transition. Additionally, phosphoproteomics and metabolomics data highlighted the importance of various phosphorylation events and metabolic pathways in EMT. Ultimately, Paul et al. were able to find two drugs that blocked EMT by inhibiting specific proteins, revealing a useful application of multiomic analyses.
Identifying biomarkers and drug targets in pancreatic cancer using multiomics
Past genomics studies of pancreatic ductal adenocarcinoma (PDAC) have uncovered mutations behind the disease, but haven’t yet led to broadly effective treatments. To improve our understanding of this devistating disease, Cao et al. conducted a multiomic analysis that compared genetic mutations with changes to the proteome and revealed new information on how proteins and associated proteoforms impact disease outcomes.
Specifically, they linked mutations in the KRAS gene to changes in specific glycoproteins that could lead to therapeutics in the future. The authors also found 12 proteins upregulated in PDAC that they suggest could be potential biomarkers for the cancer.
Proteomics aids the development of PROTAC cancer therapeutics
Targeting tumor proteins with proteolysis-targeting chimeras (PROTACs) is one way to destroy them. PROTACs bind target proteins to an ubiquitination ligase, which marks them for proteolysis. Khan et al. designed a PROTAC that targets the anti-apoptotic BCL-XL protein for destruction in leukemia cells and used a proteomic analysis to test how effective it was. Using proteomics, the team showed that only BCL-XL was degraded by the PROTAC in vitro. Later, they used the PROTAC to kill leukemia cells, while leaving non-cancerous cells minimally effected. Proteomic techniques could be transformational for this promising and creative new form of cancer treatment.
Subtyping HNSCC with a multiomic analysis
Improved understanding and subtyping of head and neck squamous cell carcinoma (HNSCC) could lead to better treatments for this challenging cancer. Huang et al. used a multiomic approach to combine data from genomics, transcriptomics, and proteomics to classify patient samples and identify clear subgroups with associated biomarkers.
Their multiomics work also highlighted that connections between DNA, mRNA and associated proteins in HNSCC are not straightforward, meaning that combining data from multiple levels of biology is necessary to understand this cancer. By pairing genomics, proteomics and more, this team identified biological pathways implicated in the disease, including some that might be targeted by existing immunotherapy drugs.
Identifying drivers and subtypes in colon cancer using proteomics
By pairing genomics, transcriptomics and proteomics, Vasaikar et al. were able to associate genes with proteins involved with colon cancer and identify new colon cancer subtypes with different underlying biologies. They examined samples with hyper-mutated DNA and those without and each displayed mutations in different sets of genes. The researchers linked genetic mutations to their associated proteins, yielding a dataset of potential drug targets. They also found several kinase-signaling pathways involved in the progression of colon cancer, as well as tumor-specific proteins that could be used in the creation of cancer vaccines in the future.
Proteomic analyses reveal new breast cancer subtypes
Breast cancer can present differently in individual patients based on their genetics, the presence of specific mutations, and other factors. To better subtype breast cancer, Asleh et al. conducted a discovery proteomics analysis of 300 samples from breast cancer patients, and grouped samples into four groups based on proteomic similarities.
Digging deeper, Asleh et al. linked certain protein profiles from their broadscale proteomic analysis to outcomes, showing that some subtypes had better prognoses than others. For example, proteins showing a strong immune response were correlated with a higher survival rate. Their breast cancer proteomics study also found 85 proteins associated with lower odds of a relapse in triple-negative breast cancer, which is more deadly.
Multiomic study highlights value of the proteome for prostate cancer prognoses
Using a multiomic approach that included genomic, transcriptomic, and proteomic analyses, Sinha et al. examined samples of prostate tumors to see how genetic alterations impact other omes. They identified new subtypes based on both genomics and proteomics, but note the two classes of subtypes didn’t align well.
Their findings highlight that genetic changes in prostate cancer are often poorly linked to changes in the proteome, meaning that other factors influence proteomic changes. That means genomics can only partly unveil the mechanics of prostate cancer, and indicates that a multiomic approach may find new biomarkers for the disease.
Check out the Translating Proteomics podcast for an exciting discussion about the applications of proteomics
MORE ARTICLES