Applications of proteomics in cancer – Revealing molecular mechanisms of the epithelial to mesenchymal transition

Tyler Ford
October 18, 2023
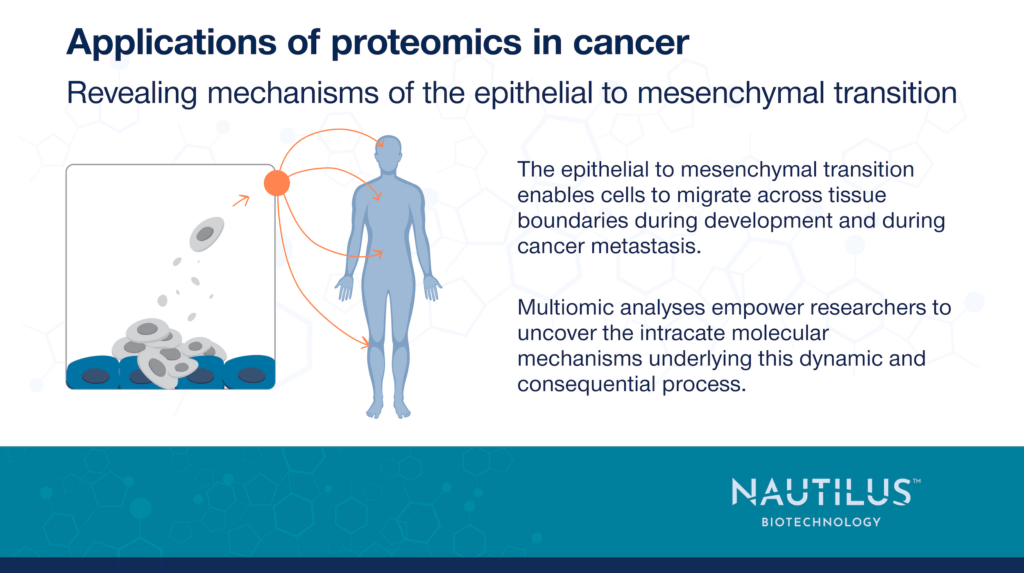
The epithelial to mesenchymal transition (EMT) is an incredibly important process in both healthy development and cancer. In development, this transition enables cells to migrate to various locations in a developing organism and seed the creation of tissues and organs. Tumor cells that undergo EMT, on the other hand, gain the ability to travel throughout the body and seed the growth of metastases.
There have been many studies of the processes underlying EMT, but omics-scale technologies are enabling researchers to move beyond studies of single proteins or pathways. These technologies empower scientists to get a holistic understanding of the many coordinated pathways underlying this complex and dynamic process and drive the identification of key points where the process can be inhibited.
In this blog post, we cover fantastic work by Paul et al. 2023 demonstrating the ways proteomics, transcriptomics, metabolomics, and more can be combined to create detailed, multiomic, and mechanistic models of complex and incredibly consequential processes like EMT.
Uncovering EMT mechanisms through multiomic analyses
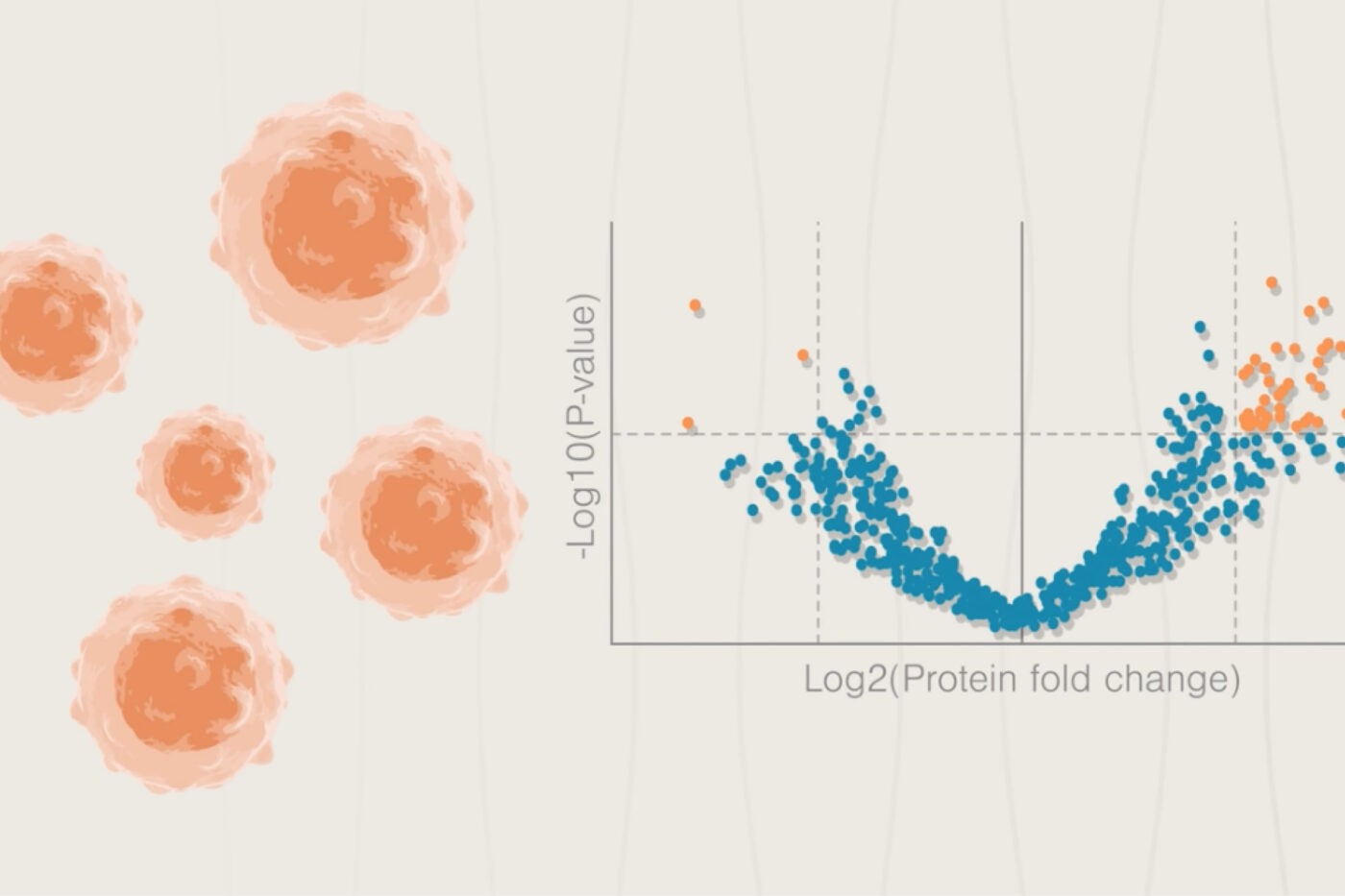
Paul et al. 2023 performed extensive studies on the MCF10A human mammary epithelial cell line in vitro. They treated the cells with the TGF-beta cytokine over a period of 12 days to induce them to undergo EMT and performed multiomic analyses including, proteomics, phosphoproteomics, transcriptomics, metabolomics, and single cell RNA sequencing at various time points. They even measured a variety of subcellular proteomes. Then, they used bioinformatics techniques to identify relationships between the various omic data sets and progression through EMT.
There was often poor correlation across the omes. For example, the coefficient of determination between mRNA and any of the other omes was no higher than 0.2 across all time points. This highlights that it is essential to do multiomic analyses to get a mechanistic understanding of complex cellular processes like EMT.
Even though the various omics layers were often poorly correlated across time points, the researchers could cluster data across the time points and partition distinct stages of EMT:
- Epithelial (Day 1)
- Transition state 1 (Days 2-3)
- Transition state 2 (Day 4)
- Mesenchymal (Days 5-12)
The multiomic data showed that distinct molecular pathways were active at these different stages. For example, pathways enriched in the mesenchymal stage included smooth muscle contraction, regulation of TNFR1, NF-kB signaling, transferrin endocytosis and recycling, and insulin receptor signaling. These results show that cells can be manipulated in different ways depending on what stage they are in and what pathways are active in that stage.
View our animation to see how next-generation proteomics can fuel cancer research
Phosphoproteomics and metabolomics serve as important indicators of protein activity
Digging deeper, the researchers used phosphoproteomics to identify highly phosphorylated peptides and metabolomics to discover metabolic pathways active at different stages of EMT. Phosphorylation was often poorly correlated and sometimes anti-correlated with protein abundance, but could be used to hypothesize roles for particular kinases and phosphorylation events. For example, the researchers hypothesized that phosphorylation of MICAL3 during EMT regulated its nuclear localization and went on to show that knock down of MICAL3 inhibited TGF-beta induced EMT.
With metabolomics, they discovered the arachidonic acid metabolism pathway was upregulated during EMT. siRNA mediated knock down of a key enzyme in this pathway prevented EMT. This is an exciting finding given that this pathway is not well-studied in EMT.
Single-cell RNA sequencing enabled the researchers to identify individual cells at various stages along the EMT trajectory. Such analyses pointed to transcription factors important at steps along the transition and helped the researchers infer intercellular communication pathways active in cells undergoing EMT.
Combining data from these omic layers enabled the researchers to generate a mechanistic model of EMT. This model identified a cascade of pathways and controller proteins driving EMT, many of which had not been identified before. It helped the researchers identify two drugs capable of blocking EMT through the inhibition of the SMO and CAMK-II proteins. Applying these inhibitors in a model of mammary cell invasiveness showed that the drugs effectively decreased invasiveness in vitro. These and similar inhibitors may have therapeutic potential in cancer.
Further insights into biological processes enabled by next-generation proteomics
The research efforts described above show just how important it is to move beyond single omic measurements and instead develop multiomic perspectives on biological processes. Had these researchers stuck to one type of measurement, they may have been misled about the activities of various pathways and would not have been able to create such a rich model to identify new drug targets.
We hope to make studies like this one accessible to a wide variety of researchers through easy-to-use, next-generation proteomics technologies. In addition, we plan to enable researchers to achieve more comprehensive assessments of the proteome through a platform that is designed to provide in-depth views of proteins across the wide dynamic range of the proteome.
We hope to give researchers the ability to develop in-depth models of broad biological processes and thereby identify means of manipulating these processes to treat disease and more.
MORE ARTICLES

