While toxicology has improved human health and safety, comprehensive proteomic analyses could reveal the molecular innerworkings associated with toxicity and off-target drug effects – improving toxicologists’ ability to predict adverse outcomes in pre-clinical models and beyond.
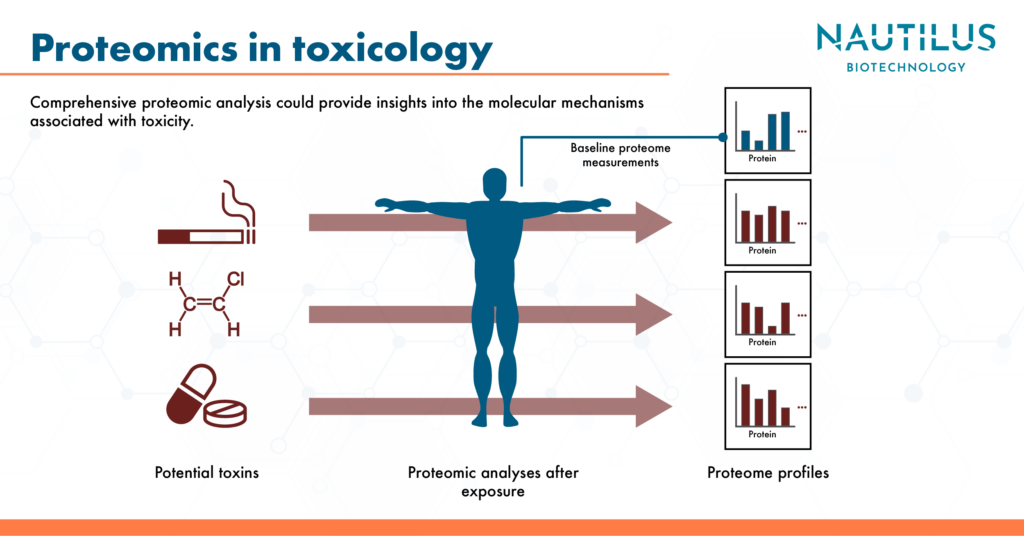
A role for comprehensive proteomics in toxicology
If you have ever drunk water from the tap, taken a dose of medication, or breathed in fresh air without accidentally poisoning yourself – you can thank toxicology. Researchers in this important field study the detection and effects of possible toxins. Toxin is a broad term and can refer to anything from harmful chemical exposures, to substances in prescription medicine. Work in this field provides us with the ability to detect harmful substances, determine the dose at which they become dangerous, and ascertain whether a substance has detrimental effects in the first place. However, current toxicology techniques are far from perfect, particularly in the area of pharmaceutical development, often missing unwanted side effects of drugs. Having new technologies that can better analyze potential toxins and their effects will go a long way toward improving human health and safety.
In this post, we will explore the potentially powerful role of comprehensive proteomic analysis in identifying biomarkers of toxicity and assessing off-target effects of drugs at the molecular level.
How do you know if something is poisoning you?
Despite what we see in the movies, ingesting toxic substances does not always lead to instantaneous foaming at the mouth or a dramatic collapse into a plate of (apparently poisoned) food. Many of the toxins we encounter on a daily basis have much more subtle impacts that accumulate over time – DNA damage, buildup of harmful agents, or worsening of preexisting conditions. A major focus of toxicology is discovering the damage and agents responsible before subtle impacts turn into chronic illnesses.
Current toxicology methods evaluate effects of substances in vitro or in animal models to uncover endpoints that are adversely impacted, such as cell death or the occurrence of developmental defects. In vitro experiments utilize cells, tissues, or organs to study toxin effects in a controlled environment – evaluating cytotoxicity, cell proliferation, membrane integrity, and other localized outcomes. The simplified nature of in vitro models often limits accurate representation of toxicity response.
In contrast, animal models provide a means for evaluating toxin distribution, effect on physiology, and biochemical interactions across several organs in a live organism. Unfortunately, the significant biological differences – such as cell distribution or blood pH – between test animals and humans reduce the predictability of these models. These techniques draw an incomplete picture and fail to reveal impacts on more nuanced molecular and biochemical pathways, especially those involved in homeostatic processes.
The vast potential of proteomics in toxicology
Transcriptomics is often used to assess toxicological response in these experiments, but mRNA transcript levels only partially correlate to protein levels due to a variety of biological processes. More accurate assessments of the ways toxins alter proteins associated with various cellular functions would provide informative measures for toxicity. For example, post translational modifications of proteins related to cell growth, division, stress, or repair– like p53 or hypoxia inducible factor – might reveal adverse effects that are not obvious from the mRNA alone.
In a 2014 review covering proteomics for systems toxicology, the authors highlighted the need for comprehensive insights into molecular toxicological mechanisms in order to improve toxicity predictions. The ability to measure changes at the molecular level within cells, tissues, and entire animals in response to exposure would allow more specific identification of relevant pathways in model systems. Understanding which molecular players are involved, and what modifications initiate their active state during a toxicity response would help to narrow in on these toxicological mechanisms.
The Nautilus Proteome Analysis Platform may be able to fill this technological gap. It is designed to rigorously probe protein abundance and modifications at varying levels of biological complexity. This detailed proteomic analysis could enhance the predictive ability of animal and in vitro toxicity models through a sensitive assessment of subtle changes in homeostatic biomolecules prior to cell death. It could also reveal the molecular basis behind toxin interactions in specific organs.
Toxicology (and proteomics) in drug discovery and safety
A related arm of toxicology focuses on the potential harm that can arise from pharmaceuticals, often known as off-target effects. We are all familiar with the long lists of side effects recited at the end of pharmaceutical commercials. These adverse side effects often arise from unintended, and unwanted, interactions with non-target molecules, pathways, or cell types. Sometimes, off-target effects are sufficient to knock drug candidates out of clinical trials, but the adverse effects of many drugs are not apparent until they’ve already hit the market because our current methods of measuring toxicity are not sufficient. Both human health and the costly drug development process would greatly benefit from a way to predict off-target effects in pre-clinical studies.
Proteomics has previously been used to expose off-target effects. An example of this was the use of mass spectrometry to uncover the molecular link between liver toxicity and Troglitazone, a since withdrawn type-II diabetes medication. Following removal from the market, researchers utilized proteomics to track Troglitazone associated mitochondrial protein changes in rodents, revealing deterioration of relevant proteins and dysfunction of transport systems leading to liver injury. Troglitazone and its associated risk of liver injury in some patients highlight the need for a robust method for detecting early signs of drug toxicity.
The Nautilus Proteome Analysis Platform could provide the expanded proteome coverage needed to reveal off-target interactions in the early stages of drug development. Rather than only investigating proteins of interest or those that are biologically linked to drug targets, robust proteome analyses can show the whole picture through broadscale analyses and targeted proteoform studies – they can clue researchers on to seemingly unrelated molecular modulations that may translate to nasty side effects down the line. Thus, with a comprehensive proteomic analysis platform designed to reveal these issues, researchers can mitigate them before a drug is released or provide physicians with the information needed to avoid or treat the toxic side effects. This may lead to safer drugs for all.
Listen the Translating Proteomics podcast for a fascinating discussion of the applications of proteomics
MORE ARTICLES