Single-molecule analyses study proteins and other target analytes at the level of individual molecules. In proteomic analyses, researchers can use single-molecule techniques to quantify proteins and proteoforms at the highest resolution possible. Proteomic analysis platforms capable of robust, scalable single-molecule analyses could revolutionize the field of biology.
When there’s a question about the inner workings of cells, biologists often turn to the proteins present, also known as the proteome. Broadscale proteomic analyses, like those that compare the proteomes of many different cells, can be incredibly enlightening when it comes to understanding the big picture. They can reveal what biological pathways are active in different cells and point researchers to the molecular basis of a biological function. These proteomic analyses have proven invaluable — but traditional techniques often measure proteins in bulk and don’t provide insights into single protein molecules.
Excitingly, we are poised for a single-molecule proteomics revolution with the development of next-generation proteomics technologies like the NautilusTM Proteome Analysis Platform that aim to achieve single-molecule analysis of all proteins in a sample. These analyses could reveal a sample’s proteome at depths never before possible, including quantifying proteins present in very low abundance and identifying protein variants known as proteoforms. Each proteoform is defined by the full set of modifications it has, whether those modifications come from alternative splicing, post-translational modification, or any other source. Thus identifying proteoforms requires single-molecule analysis of full-length proteins.
Read our preprint to explore novel proteoform data and its potential in drug development
Critically, we’re designing a platform capable of single-molecule analysis of intact proteins. Many other technologies only analyze small pieces of proteins called peptides at the single-molecule level. This makes it difficult to determine which peptides came from the same full-length protein and obscures the many possible combinations of modifications present on the full protein. Overall, without the ability to analyze intact proteins at the single-molecule level, it is very difficult to identify and quantify the precise proteins and proteoforms present.
Powered by a platforms like ours, single-molecule protein studies can reveal the identity of proteins across the dynamic range of substantively the entire proteome in a sample. That knowledge could play an important role in functional proteomics studies that attempt to reveal the roles specific proteins play in the body, whether that’s catalyzing chemical reactions or allowing a pathogen to enter our cells. Furthermore, single-molecule analysis is critical for targeted proteomics, as it enables researchers to elucidate the precise proteoforms present in a sample.
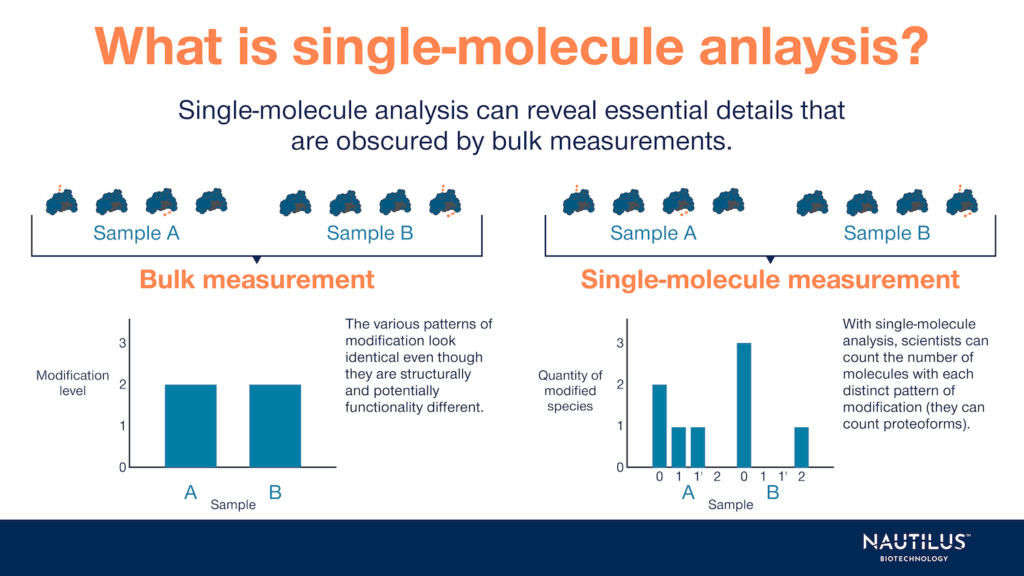
In general, single-molecule analyses can reveal mechanistic details that are obscured by bulk measurements, and single-molecule techniques are useful across a wide variety of fields and applications. Here’s a quick snapshot of when single-molecule analysis is most helpful — in proteomics and beyond — and what researchers can learn from it.
Watch this animation to learn how single-molecule analysis enables proteoform studies on the Nautilus Platform:
Examples of single-molecule analysis
Single-molecule measurements give the highest resolution and highest sensitivity possible when quantifying proteins or other molecules present in a sample. A single-molecule analysis can reveal even tiny variations to proteins or other molecules.
This sort of resolution into the proteome or other types of molecules can be necessary for diagnosing disease, understanding the state of tissues, or finding treatments for disease. For instance, in a recent report in Nature Biotechnology, researchers in Israel developed a single-molecule assay that detects and quantifies protein complexes called nucleosomes that have been modified in various ways. The assay specifically measures these molecules in blood plasma and may enable researchers to use the modified nucleosomes as biomarkers for the early detection of cancer.
Single-molecule analysis can also enable scientists to look at the mechanics of proteins involved in various pathways or reactions, since individual protein molecules can change over the course of a reaction. Without single-molecule techniques, it would be impossible to make out the precise changes related to the mechanisms of these reactions — lower resolution analyses would instead give a blurry, averaged-over-time picture of the various steps and stages.
Many such single-molecule analysis techniques are not proteomics techniques, but still provide high resolution insights into protein function. For example, a technique called single-molecule force spectroscopy is increasingly used to study the dynamics of folding and unfolding proteins. In a recent review in Langmuir, a journal of the American Chemical Society, the author shows how this type of single-molecule analysis can be used to investigate various proteins like calcium-binding proteins, which are fundamental to cellular communication. Likewise, researchers in a recent study in Nucleic Acids Research used single-molecule analysis to characterize protein-DNA interactions like those that occur during the repair of damaged DNA.
Of course, single-molecule measurements aren’t always necessary and could be overkill for certain situations. For example, if there’s a protein biomarker associated with a disease, a broad look may be all that’s needed to show whether the biomarker’s abundance is elevated (or reduced) in a sample. In other cases, including early detection of some disease biomarkers, the proteins are in low enough abundance to require single-molecule sensitivity.
Single-molecule analysis with the Nautilus Platform
We’re developing the Nautilus Platform to enable more accessible, efficient, and sensitive proteomics. This next-generation proteomics technology is designed to make use of high-throughput, single-molecule analysis that will supercharge discovery. Learn more about our technology and the steps we’re taking to design the ideal proteomic analysis platform.
MORE ARTICLES


