Nautilus at HUPO 2025 – Redefining proteomics with single-molecule Iterative Mapping
Nautilus Biotechnology
November 21, 2025

The Nautilus team just returned from HUPO 2025 in snowy Toronto, where the convention center buzzed with presentations, posters, and conversations about the future of proteomics. Scientists shared insights on how the proteome changes in health and disease, and how new tools offer varying breadth and detail in their analyses.
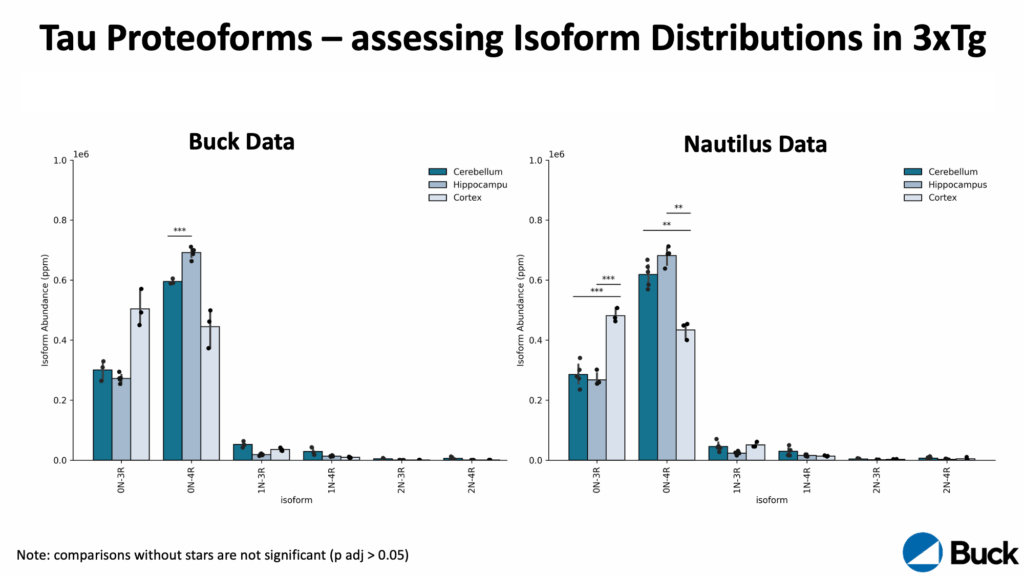
Our focus was clear: showing how the Nautilus Platform uses Iterative Mapping to eliminate the classic tradeoff between breadth (proteins detected) and detail (information per protein). We were proud to host Professor Birgit Schilling from the Buck Institute for Research on Aging, who presented how her lab is using the Nautilus Platform to quantify tau proteoform groups across brain regions in a mouse model of Alzheimer’s disease. This was a huge milestone for us—the first time a researcher shared novel data generated on a Nautilus instrument in their own lab.
Access the seminar here and register for our January 14, 2026 Select Science webinar to see new data and ask questions directly to Professor Schilling, Nautilus co-founder and Chief Scientist Parag Mallick, and Nautilus VP of Scientific Engagement Sheri Wilcox.
Professor Birgit Schilling presents data generated on the Nautilus Platform
Just before HUPO 2025, we revealed that Professor Schilling has a field evaluation unit in her lab. At HUPO 2025, she shared preliminary data demonstrating the reproducible quantification of tau proteoforms across brain regions in a mouse model of Alzheimer’s disease on that instrument. Her work showcases the quality of the data generated on the Nautilus Platform and begins to reveal differences in tau proteoform landscapes across brain regions. Understanding these landscapes may be critical to unlocking how tau impacts aging and cognitive decline.
In her presentation, Birgit stressed how thrilled she was to see this novel single-molecule data and presented a variety of preliminary hypotheses surrounding the proteoform variability she observed. Birgit’s excitement around these hypotheses was contagious, and we cannot wait to see what she uncovers in future work.
This presentation was also a critical demonstration of how the Nautilus Platform provides unprecedented breadth in terms of the quantification of hundreds of tau proteoforms and detail in terms of sensitive, precise, and reproducible single-molecule proteoform quantification. It is inspiring to see that even this preliminary data is stimulating the development of many new hypotheses – the Nautilus Platform is designed to open exactly these kinds of new research avenues.
As you’ll hear if you access the recording of the seminar, the audience Q&A showed the HUPO community is impressed with our progress. Some select quotes include:
- “Great, amazing work. Very, very impressive.”
- “Congratulations to the whole team because you’ve made amazing progress.”
- “My heart unironically started beating faster.”
Many in the HUPO community later remarked that they saw this seminar as a turning point for us. Thanks to all who attended our seminar and made this HUPO a special one!
The need to redefine proteomics
Proteomics has advanced in detecting more proteins, but researchers want more than detection—they want meaning. A common theme at HUPO was that different technologies detect different proteomes because they identify proteins with limited information: a few peptides for mass spectrometry or a couple epitopes for affinity-based methods. These partial views often don’t overlap, don’t provide a lot of information about each protein detected, and may represent different isoforms or proteoforms. To compensate, researchers combine multiple technologies and do extensive follow-up work to interpret what the differences mean.
Our takeaway was that proteomics research can be accelerated through a technology like ours that provides breadth and detail by measuring all the proteins in the proteome while providing as much information as possible about each protein molecule quantified.
A drive toward understanding relationships between proteins and function
The reason researchers want breadth and detail in their proteomic measurements is they want to understand how proteins impact function. They are not satisfied with seeing that a protein is significantly upregulated in one cell versus another, they want to know the degree to which it’s upregulated and what it’s doing. Researchers answer the question “What is this protein doing?” with follow-up experiments involving interaction proteomics, genetic knockouts, targeted analysis of posttranslational modifications, and more.
It’s exciting that researchers are beginning to use our platform when this focus on function is so strong. Our platform is designed to show not just single-molecule differences in protein quantity, but also how the makeup of each individual protein molecule changes. This is critical because it is through changes in molecular composition that researchers assess things like protein structure and interactions, both of which are key to function. For instance, through its proteoform analysis capabilities, our platform can show how the combinations of posttranslational modifications on single protein molecules change. Researchers should be able to model how these different combinations alter protein structure and activity at scale. In addition, many methods for assessing protein-protein interactions involve the addition of tags to molecules that a target protein interacts with. Our platform should be well-suited to detecting such tags in future analyses. The key is that single-molecule analysis is well-aligned with functional analysis.
HUPO 2025 left us more excited than ever to see how researchers will use our platform to develop a thorough understanding of biological function at scale.
If you’d like to be one of the first to use the Nautilus Platform in your work, please reach out!
MORE ARTICLES